33 N Vanuatu#
Time-dependent implementation#
Preamble#
We are going to run this notebook in parallel using ipyparallel. To do this we launch a parallel engine with nprocs processes. Following this, all cells we wish to run in parallel need to start with the jupyter magic %%px.
%%px
Cells without the %%px magic will run locally and will not have access to anything loaded on the parallel engine.
import ipyparallel as ipp
nprocs = 2
rc = ipp.Cluster(engine_launcher_class="mpi", n=nprocs).start_and_connect_sync()
Starting 2 engines with <class 'ipyparallel.cluster.launcher.MPIEngineSetLauncher'>
Set some path information.
%%px
import sys, os, shutil
basedir = ''
if "__file__" in globals(): basedir = os.path.dirname(__file__)
sys.path.insert(0, os.path.join(basedir, os.path.pardir, os.path.pardir, os.path.pardir, 'python'))
Loading everything we need from sz_problem and also set our default plotting and output preferences.
%%px
import fenics_sz.utils
from fenics_sz.sz_problems.sz_params import allsz_params
from fenics_sz.sz_problems.sz_slab import create_slab, plot_slab
from fenics_sz.sz_problems.sz_geometry import create_sz_geometry
from fenics_sz.sz_problems.sz_tdep_dislcreep import TDDislSubductionProblem
import numpy as np
import dolfinx as df
import pyvista as pv
import pathlib
output_folder = pathlib.Path(os.path.join(basedir, "output"))
output_folder.mkdir(exist_ok=True, parents=True)
import hashlib
import zipfile
import requests
from mpi4py import MPI
comm = MPI.COMM_WORLD
my_rank = comm.rank
Parameters#
We first select the name and resolution scale, resscale and target Courant number cfl of the model.
Resolution
By default the resolution (both spatial and temporal) is low to allow for a quick runtime and smaller website size. If sufficient computational resources are available set a lower resscale and a lower cfl to get higher spatial and temporal resolutions respectively. This is necessary to get results with sufficient accuracy for scientific interpretation.
%%px
name = "33_N_Vanuatu"
resscale = 3.0
cfl = 3.0
Then load the remaining parameters from the global suite.
%%px
szdict = allsz_params[name]
if my_rank == 0:
print("{}:".format(name))
print("{:<20} {:<10}".format('Key','Value'))
print("-"*85)
for k, v in allsz_params[name].items():
if v is not None and k not in ['z0', 'z15']: print("{:<20} {}".format(k, v))
[stdout:0] 33_N_Vanuatu:
Key Value
-------------------------------------------------------------------------------------
coast_distance 70
extra_width 20
lc_depth 30
io_depth 154
dirname 33_N_Vanuatu
As 20.0
Ac 25.0
A 44
sztype oceanic
Vs 51.8
xs [0, 57.0, 86.0, 109.0, 112.5, 113.4, 119.0, 127.0, 164.0]
ys [-6, -15.0, -30.0, -70.0, -80.0, -82.5, -100.0, -125.0, -240.0]
Any of these can be modified in the dictionary.
Several additional parameters can be modified, for details see the documentation for the TDDislSubductionProblem class.
%%px
if my_rank == 0: help(TDDislSubductionProblem.__init__)
[stdout:0] Help on function __init__ in module fenics_sz.sz_problems.sz_base:
__init__(self, geom, **kwargs)
Initialize a BaseSubductionProblem.
Arguments:
* geom - an instance of a subduction zone geometry
Keyword Arguments:
required:
* A - age of subducting slab (in Myr) [required]
* Vs - incoming slab speed (in mm/yr) [required]
* sztype - type of subduction zone (either 'continental' or 'oceanic') [required]
* Ac - age of the overriding plate (in Myr) [required if sztype is 'oceanic']
* As - age of subduction (in Myr) [required if sztype is 'oceanic']
* qs - surface heat flux (in W/m^2) [required if sztype is 'continental']
optional:
* Ts - surface temperature (deg C, corresponds to non-dim)
* Tm - mantle temperature (deg C, corresponds to non-dim)
* kc - crustal thermal conductivity (non-dim) [only has an effect if sztype is 'continental']
* km - mantle thermal conductivity (non-dim)
* rhoc - crustal density (non-dim) [only has an effect if sztype is 'continental']
* rhom - mantle density (non-dim)
* cp - isobaric heat capacity (non-dim)
* H1 - upper crustal volumetric heat production (non-dim) [only has an effect if sztype is 'continental']
* H2 - lower crustal volumetric heat production (non-dim) [only has an effect if sztype is 'continental']
optional (dislocation creep rheology):
* etamax - maximum viscosity (Pas) [only relevant for dislocation creep rheologies]
* nsigma - stress viscosity power law exponents (non-dim) [only relevant for dislocation creep rheologies]
* Aeta - pre-exponential viscosity constant (Pa s^(1/n)) [only relevant for dislocation creep rheologies]
* E - viscosity activation energy (J/mol) [only relevant for dislocation creep rheologies]
Setup#
Setup a slab.
%%px
slab = create_slab(szdict['xs'], szdict['ys'], resscale, szdict['lc_depth'])
if my_rank == 0: _ = plot_slab(slab)
[output:0]
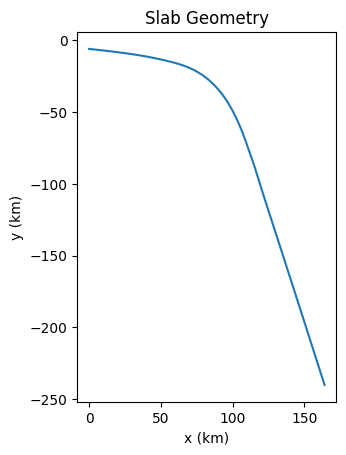
Create the subduction zome geometry around the slab.
%%px
geom = create_sz_geometry(slab, resscale, szdict['sztype'], szdict['io_depth'], szdict['extra_width'],
szdict['coast_distance'], szdict['lc_depth'], szdict['uc_depth'])
if my_rank == 0: _ = geom.plot()
[output:0]
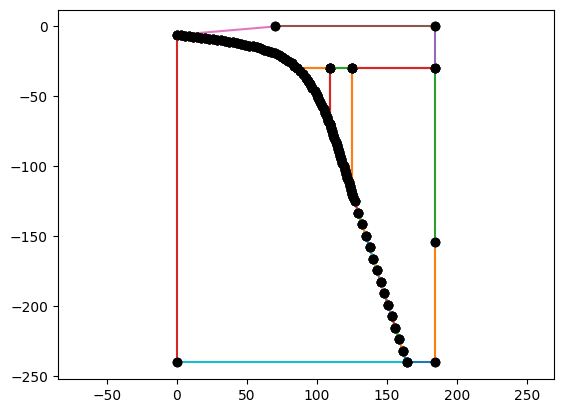
Finally, declare the TDDislSubductionProblem problem class using the dictionary of parameters.
%%px
sz = TDDislSubductionProblem(geom, **szdict)
Solve#
Solve using a dislocation creep rheology.
%%px
# Select the timestep based on the approximate target Courant number
dt = cfl*resscale/szdict['Vs']
# Reduce the timestep to get an integer number of timesteps
dt = szdict['As']/np.ceil(szdict['As']/dt)
sz.solve(szdict['As'], dt, theta=0.5, rtol=1.e-1, verbosity=1)
[stdout:0] Entering timeloop with 116 steps (dt = 0.172414 Myr, final time = 20 Myr)
Finished timeloop after 116 steps (final time = 20 Myr)
Plot#
Plot the solution.
%%px
plotter = pv.Plotter()
fenics_sz.utils.plot.plot_scalar(sz.T_i, plotter=plotter, scale=sz.T0, gather=True, cmap='coolwarm', scalar_bar_args={'title': 'Temperature (deg C)', 'bold':True})
fenics_sz.utils.plot.plot_vector_glyphs(sz.vw_i, plotter=plotter, gather=True, factor=0.1, color='k', scale=fenics_sz.utils.mps_to_mmpyr(sz.v0))
fenics_sz.utils.plot.plot_vector_glyphs(sz.vs_i, plotter=plotter, gather=True, factor=0.1, color='k', scale=fenics_sz.utils.mps_to_mmpyr(sz.v0))
geom.pyvistaplot(plotter=plotter, color='green', width=2)
cdpt = slab.findpoint('Slab::FullCouplingDepth')
fenics_sz.utils.plot.plot_points([[cdpt.x, cdpt.y, 0.0]], plotter=plotter, render_points_as_spheres=True, point_size=10.0, color='green')
if my_rank == 0:
fenics_sz.utils.plot.plot_show(plotter)
fenics_sz.utils.plot.plot_save(plotter, output_folder / "{}_td_solution_resscale_{:.2f}_cfl_{:.2f}.png".format(name, resscale, cfl,))
[stderr:1] 2026-04-30 05:06:40.159 ( 14.716s) [ 7F351A616140]vtkXOpenGLRenderWindow.:1416 WARN| bad X server connection. DISPLAY=:99
[stderr:0] 2026-04-30 05:06:40.155 ( 14.712s) [ 7FAEEE7BA140]vtkXOpenGLRenderWindow.:1416 WARN| bad X server connection. DISPLAY=:99
[output:0]
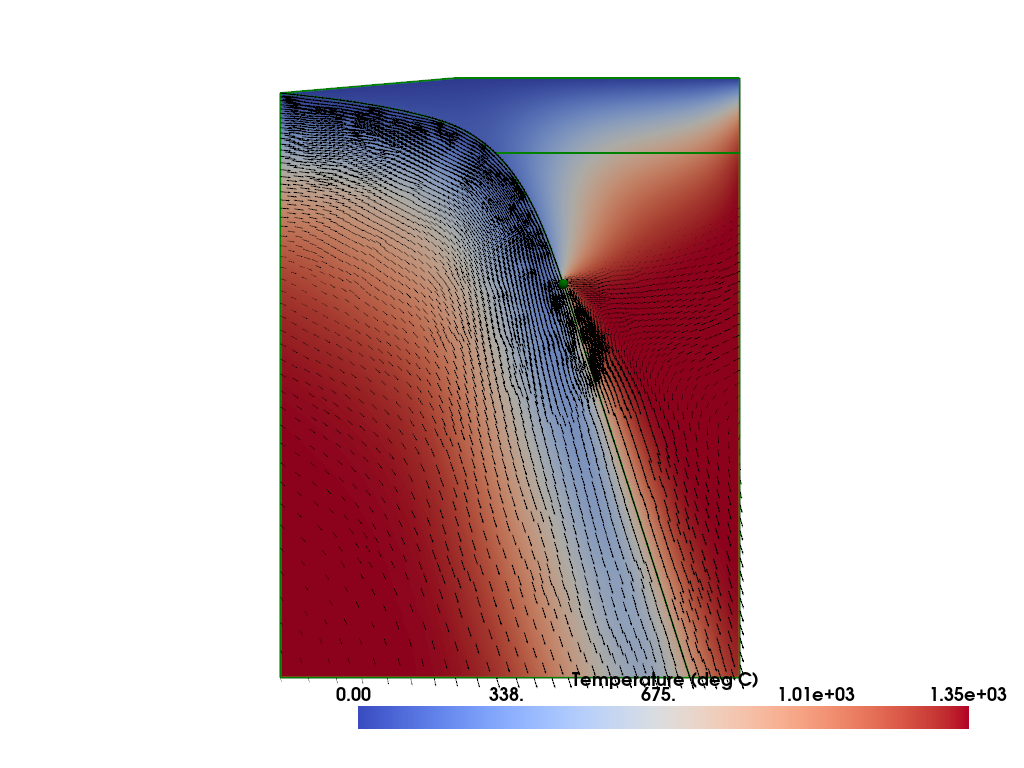
Save it to disk so that it can be examined with other visualization software (e.g. Paraview).
%%px
filename = output_folder / "{}_td_solution_resscale_{:.2f}_cfl_{:.2f}.bp".format(name, resscale, cfl,)
with df.io.VTXWriter(sz.mesh.comm, filename, [sz.T_i, sz.vs_i, sz.vw_i]) as vtx:
vtx.write(0.0)
# zip the .bp folder so that it can be downloaded from jupyter lab
if my_rank == 0: shutil.make_archive(str(filename), 'zip', root_dir=str(filename.parent), base_dir=str(filename.name))
Comparison#
Compare to the published result from Wilson & van Keken, PEPS, 2023 (II) and van Keken & Wilson, PEPS, 2023 (III). The original models used in these papers are also available as open-source repositories on github and zenodo.
First download the minimal necessary data from zenodo and check it is the right version.
%%px
zipfilename = pathlib.Path(os.path.join(basedir, os.path.pardir, os.path.pardir, os.path.pardir, "data", "vankeken_wilson_peps_2023_TF_lowres_minimal.zip"))
# only one process should download the data
if my_rank == 0:
if not zipfilename.is_file():
zipfileurl = 'https://zenodo.org/records/13234021/files/vankeken_wilson_peps_2023_TF_lowres_minimal.zip'
r = requests.get(zipfileurl, allow_redirects=True)
open(zipfilename, 'wb').write(r.content)
# wait until rank 0 has downloaded the data
comm.barrier()
assert hashlib.md5(open(zipfilename, 'rb').read()).hexdigest() == 'a8eca6220f9bee091e41a680d502fe0d'
%%px
tffilename = os.path.join('vankeken_wilson_peps_2023_TF_lowres_minimal', 'sz_suite_td', szdict['dirname']+'_minres_2.00_cfl_2.00.vtu')
tffilepath = os.path.join(basedir, os.path.pardir, os.path.pardir, os.path.pardir, 'data')
# one process should extract the file
if my_rank == 0:
with zipfile.ZipFile(zipfilename, 'r') as z:
z.extract(tffilename, path=tffilepath)
# other processes should wait
comm.barrier()
%%px
fxgrid = fenics_sz.utils.plot.grids_scalar(sz.T_i)[0]
tfgrid = pv.get_reader(os.path.join(tffilepath, tffilename)).read()
diffgrid = fenics_sz.utils.plot.pv_diff(fxgrid, tfgrid, field_name_map={'T':'Temperature::PotentialTemperature'}, pass_point_data=True)
%%px
# first gather the data onto rank 0
diffgrid_g = fenics_sz.utils.plot.pyvista_grids(diffgrid.cells, diffgrid.celltypes, diffgrid.points, comm, gather=True)
T_g = comm.gather(diffgrid.point_data['T'], root=0)
for r, grid in enumerate(diffgrid_g):
grid.point_data['T'] = T_g[r]
grid.set_active_scalars('T')
grid.clean(tolerance=1.e-2)
diffgrid_g
# then plot it
plotter_diff = pv.Plotter()
clim = None
for grid in diffgrid_g:
plotter_diff.add_mesh(grid, cmap='coolwarm', clim=clim, scalar_bar_args={'title': 'Temperature Difference (deg C)', 'bold':True})
geom.pyvistaplot(plotter=plotter_diff, color='green', width=2)
cdpt = slab.findpoint('Slab::FullCouplingDepth')
#fenics_sz.utils.plot.plot_points([[cdpt.x, cdpt.y, 0.0]], plotter=plotter_diff, render_points_as_spheres=True, point_size=10.0, color='green')
plotter_diff.enable_parallel_projection()
plotter_diff.view_xy()
if my_rank == 0: plotter_diff.show()
[output:0]
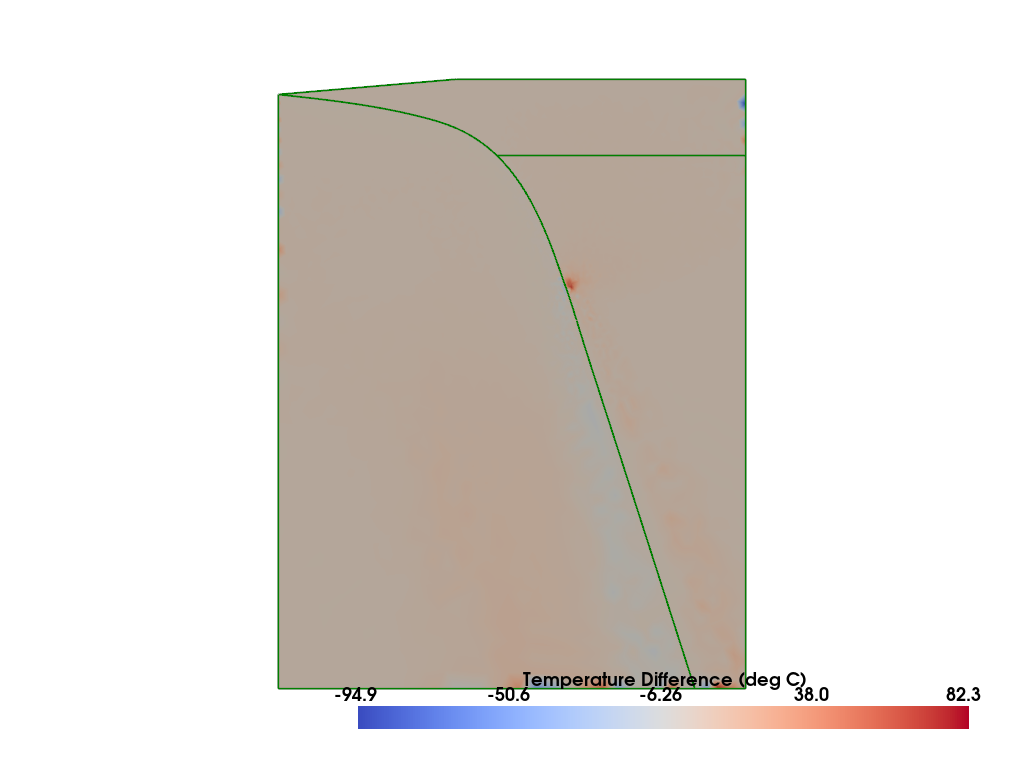
%%px
integrated_data = diffgrid.integrate_data()
totalarea = comm.allreduce(integrated_data['Area'][0], op=MPI.SUM)
error = comm.allreduce(integrated_data['T'][0], op=MPI.SUM)/totalarea
if my_rank == 0: print("Average error = {}".format(error,))
assert np.abs(error) < 5
[stdout:0] Average error = 1.0321032803704187
Shutdown#
Shutdown the ipyparallel cluster engine. Running the cell below will prevent earlier cells (with %%px as the first line) from being re-run.
rc.shutdown()
